Vi-BIO
Visual Intelligence for Biology
OmiX Microbiome Analysis Platform Prototype

OmiX Design Concepts
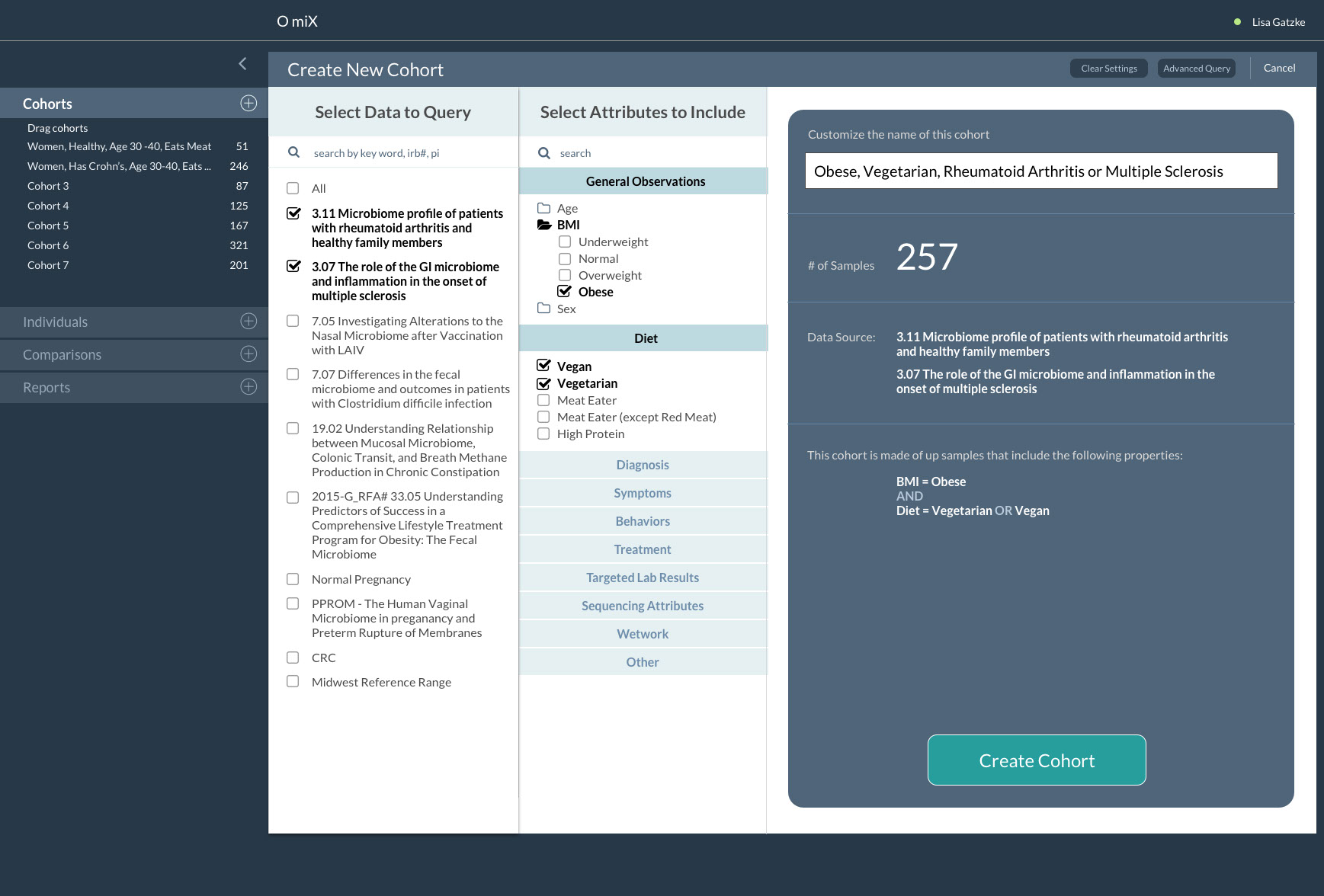
OmiX: Create a New Cohort Concept
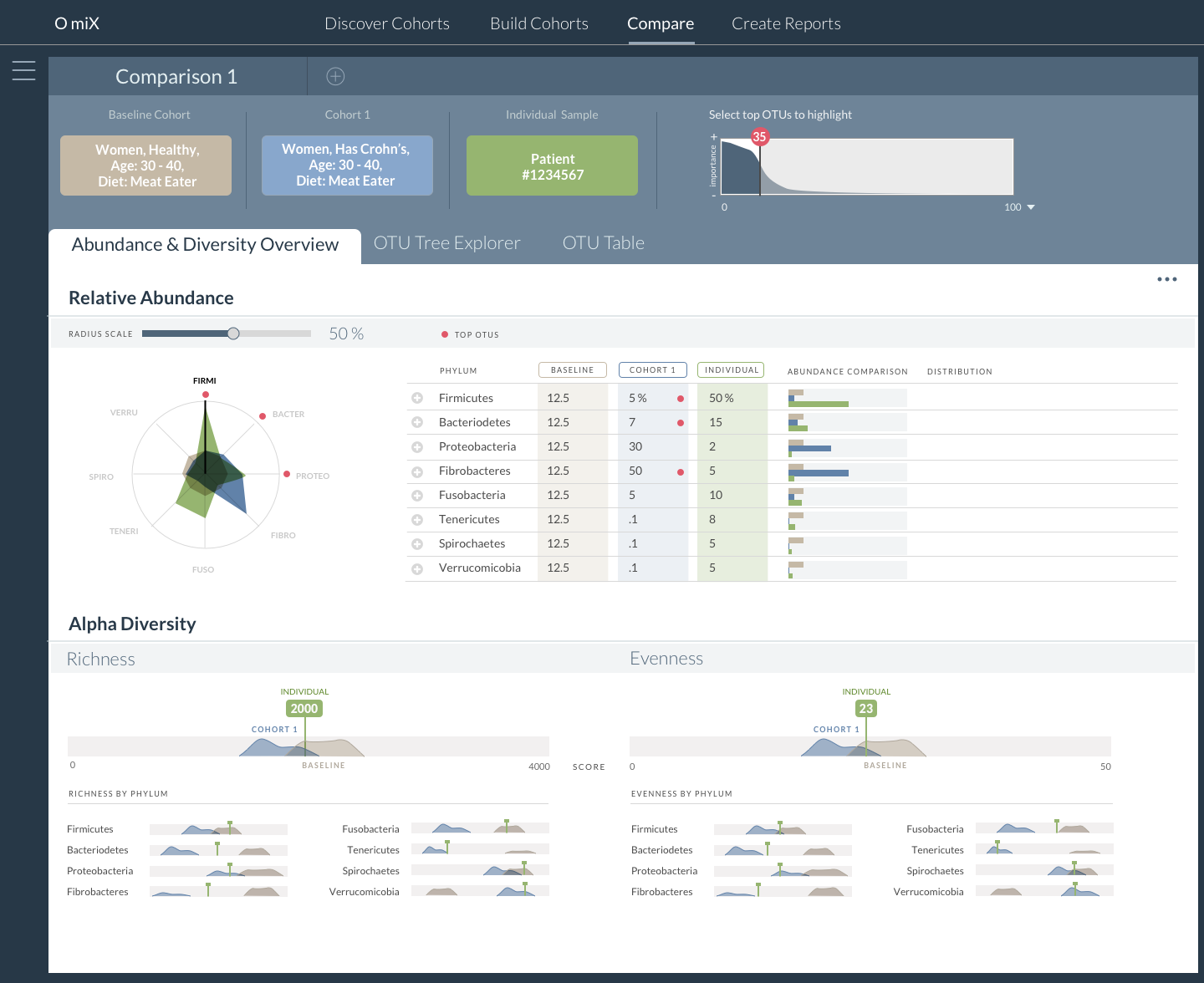
OmiX: Cohort Comparison of Microbial Abundance and Diversity
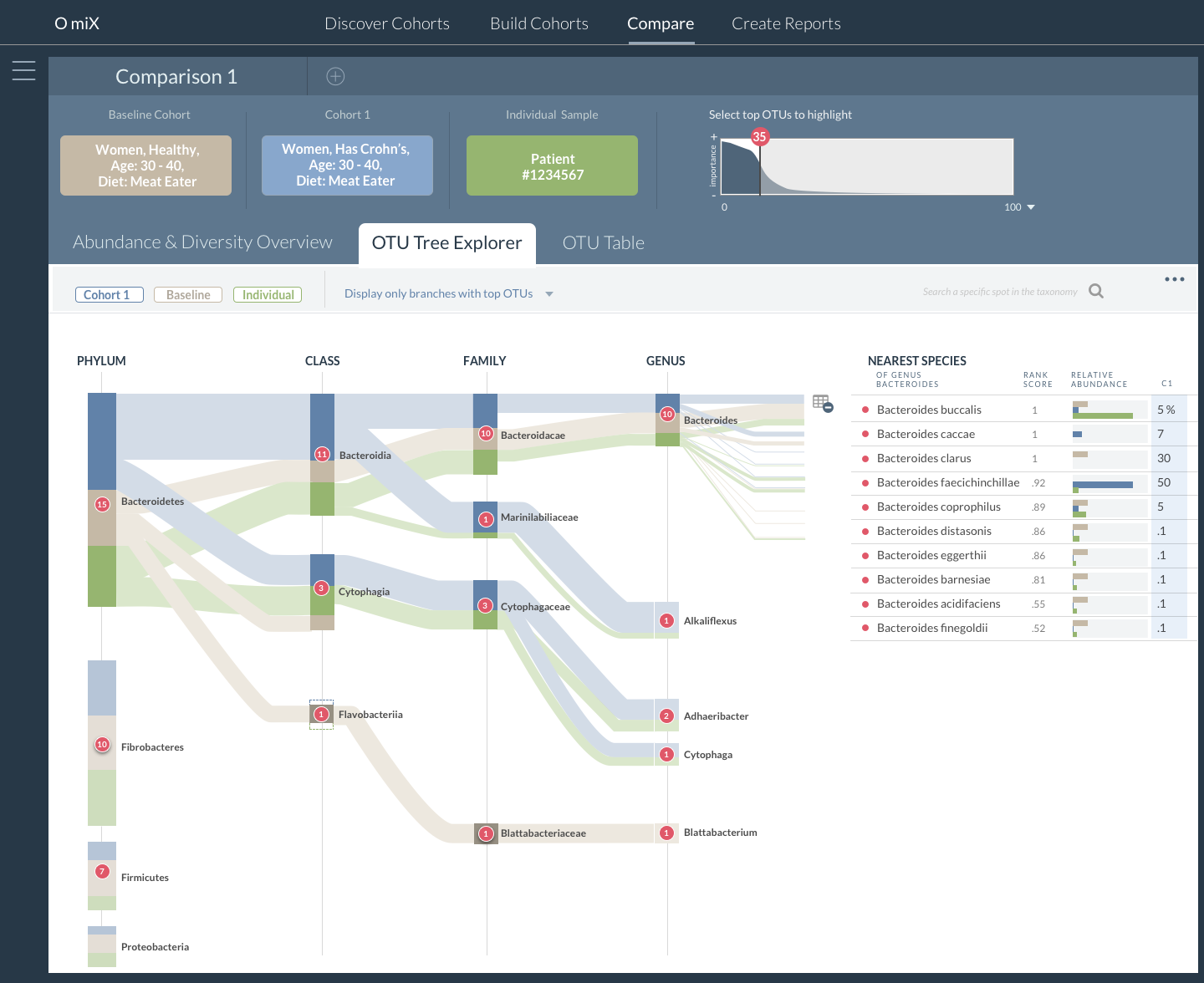
OmiX: Cohort Comparison at All Taxogenomic Levels Using the OTU Tree Explorer Visualization
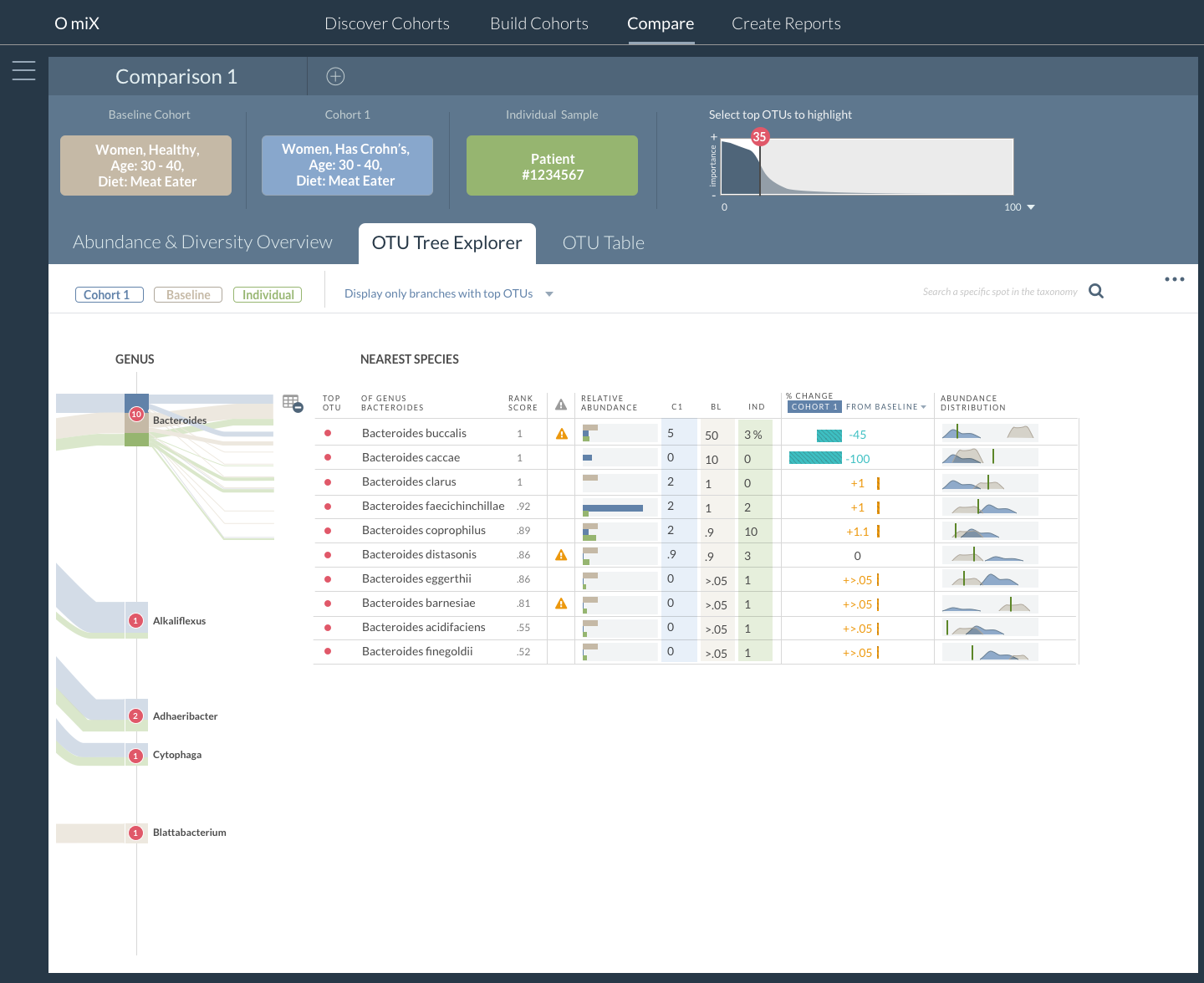
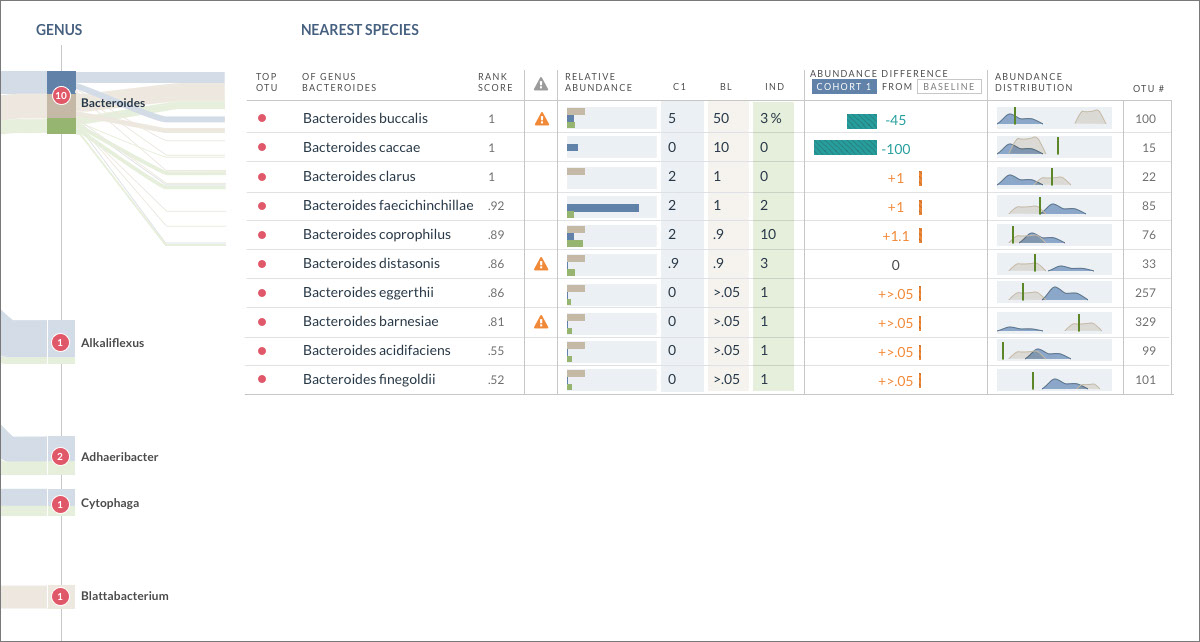
OmiX: OTU/Species Data Comparison Table
COLLABORATORS
Heidi Nelson, MD, Department of Surgery
Nick Chia, PhD., Mayo Clinic Center for Individualized Medicine
PROJECT DATES
September 2015 - December 2016
DEMO ACCESS
A working demo of the prototype of the software is now available. To request access, email support@visari.org
WHITE PAPER
Read the white paper titled, OmiXschema and Durable Core
The OmiX microbiome analysis tool is being developed in collaboration with our colleagues at the Mayo Clinic’s Center for Individualized Medicine. This project involved the design and creation of a software system that ingests microbiome data from multiple studies, processes the data, performs analytics and displays results in an interactive web-based environment.
With this tool researchers are able to effectively visualize and study the microbial populations of healthy individuals and patients with various diseases including colon cancer, rheumatoid arthritis and intestinal infections. Data about specific patients is visually portrayed in the context of other cohorts – a healthy cohort, and a cohort of people with the same disease. This method allows the viewer to see the distribution range of others so they can more clearly understand how their patient compares to others. Simply comparing a patient to an average “normal” patient is misleading when there is wide variation in what constitutes “normal”.
Important microbes are identified by the analysis and displayed on an interactive phylogenetic tree viewer to help researchers understand the relationships between key bacteria. The intention of the project is to bring microbiome research closer to clinical practice.
© Copyright 2016 University of Illinois Board of Trustees
VI-BIO - Visual Intelligence for Biology
NCSA | UNIVERSITY OF ILLINOIS
CONTACT: cbushell@illinois.edu
